Knauss legislative fellowships in Congress help build careers — and they're fun and educational. See our video and fact sheet for details.
Can computer models help restore Maryland’s oyster population? It looks like yes.
Oysters are crucial to a healthy Chesapeake Bay. They filter water, attract biodiversity and provide essential habitat for other important Bay organisms. Today, less than one percent of the oyster biomass that existed before European settlers arrived remains in the Chesapeake. So, science is giving nature a boost by cultivating oysters at the Horn Point Laboratory Oyster Hatchery in Maryland, planting them in designated rivers and creeks throughout the Bay, and monitoring their growth and abundances.
One of the focal points of these efforts in Maryland is Harris Creek, a tributary of the Choptank River, and one of the largest oyster restoration projects in the world.
While restoring oysters may seem simple — plant them and they will grow, right? — restoring oyster populations in the Chesapeake Bay requires a number of sophisticated monitoring techniques to ensure that these oyster populations are healthy and resilient.
My job is to look at the DNA of these oysters and measure genetic diversity in oysters from the hatchery as well as in oysters from restoration sites in Harris Creek to understand how it changes over time. Monitoring the genetic diversity in these oysters gives researchers an idea of their overall health. If these oysters have a lot of genetic diversity, then they are more likely to survive disease outbreaks, and adapt to changing environments. These metrics are not just measuring sticks; they are important for guiding decisions related to oyster restoration. And while I am able to measure genetic diversity at restoration sites, I am unable to measure how it may potentially change over many generations as a result of specific pressures like disease. Therefore, we model.
Computer simulations (models) are extremely useful tools for mapping natural systems in a wide range of fields including economics, social sciences, biology, and chemistry in order for researchers to gain insight into the operation of these systems. We also rely on many models in our everyday life, like those that provide weather forecasts. Natural systems are extremely complex, and understanding how they are affected as a result of various interacting processes, like climate change and/or overfishing, can be extremely difficult to measure in the field. In the past, scientists would undertake detailed field work to better understand how these systems work. That often takes a lot of effort, a lot of time, and sometimes a lot of money, not to mention this research can only explain the current system, not future conditions. However, this work was not done in vain. There are copious amounts of data from field scientists that have been recorded over many decades that researchers can use to understand natural systems.
In order to predict or better model interactions in natural systems, scientists use computers and mathematics to build models that represent these systems, much like a model train set is a scaled down representation of a real train. So, I am in the process of building a model to better understand and predict the genetic impact of hatchery supplementation within the Harris Creek Sanctuary, as well as within the broader realm of shellfish restoration programs (that supplement with hatchery produced oysters) in other parts of the world.
Similar but simpler models have been used to understand the genetic impact of salmon supplementation, but these models fail to capture the complex life history of oysters, as they were built for salmon, which are a relatively short-lived fish and spawn once and then die. Oysters, on the other hand, are a relatively long-lived species and spawn several times throughout their life and thus have overlapping generations. We wouldn’t use a model train set as a scaled down representation for an airplane, so I have to build my own scaled-down oyster model, since the salmon models don’t represent it well. Therefore, this oyster model will capture oyster life-history components such as those mentioned above, along with other important information like wild abundances before oysters were supplemented as well as the number of oysters supplemented.
I’d like to think of myself as relatively skilled in the R statistical language, which is a program researchers use to run statistics and create figures and graphs, and is what I will be using to build my oyster model; however, throughout this process, I quickly learned that there was sufficient room to improve. The amount of information on this modeling program is limited, so I was left to read the manual, which essentially consists of some very undetailed information about specific types of things I can do within the program. I remember when I finally found a report that used this program and published their code (which was a study about whales and is in no way similar to what I am doing). I simply copied their code, tried to run it, and attempted understand what was even going on. Then, I tried to edit their code to match the model I was trying to create, with extreme confusion and a lot of errors.
After carefully reading the manual (over and over) and essentially all of the available information about this program that I could get my hands on, and diligently working with the program a little bit at least three times per week for a few months, I can finally say I understand the program (and can explain it to my advisor in a way that he can understand). I can confidently describe (and build!) the oyster model that I will be creating for my dissertation. In addition to working several times a week within the program, I reached out to the program developer as well as a few others who have used the program for their research to ask for specific advice.
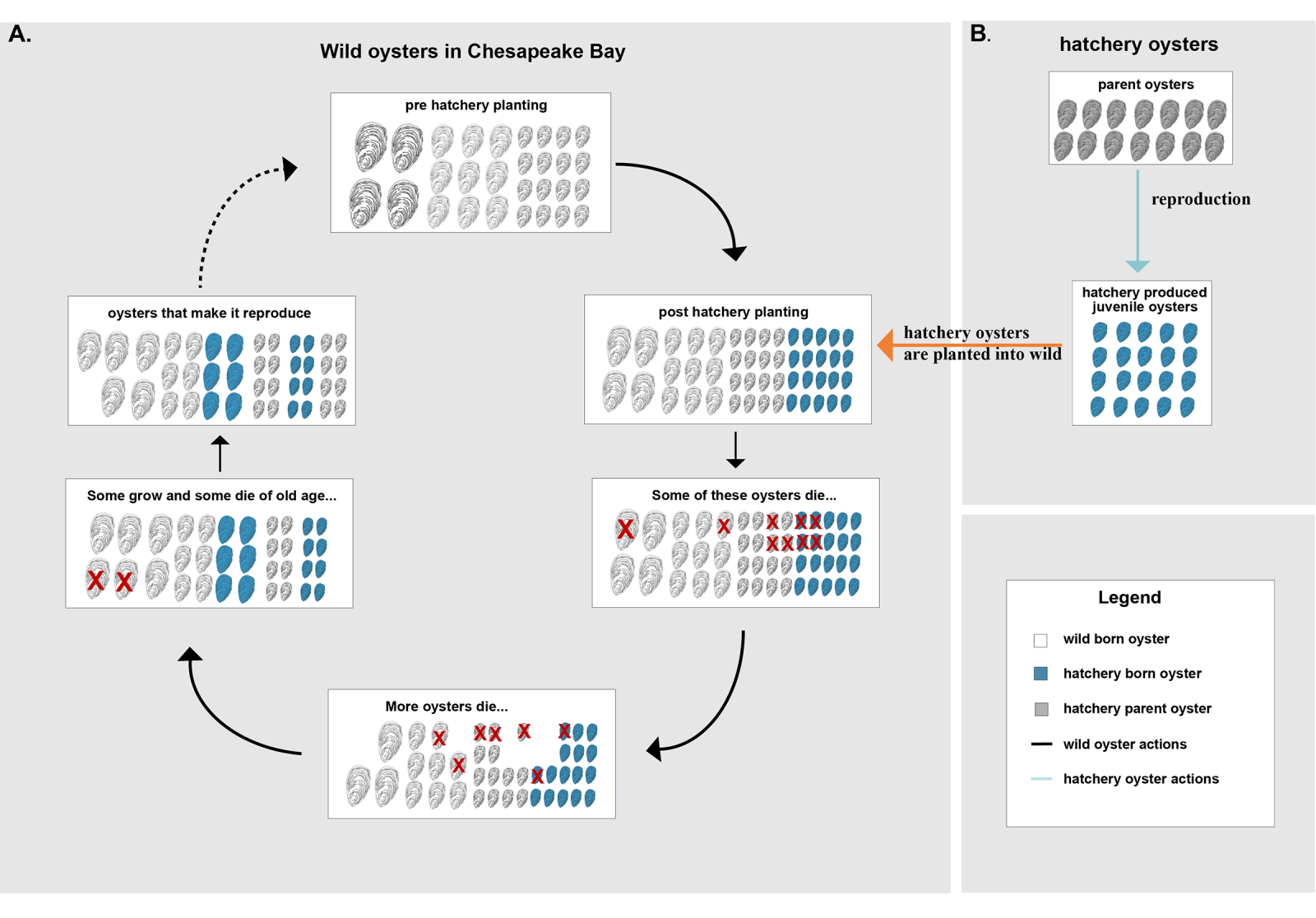
Schematic representation of the oyster model. Oysters in the wild population will be of three different ages — juveniles, medium-sized oysters, and large-sized oysters. They will go through the following steps: (A) death due to overcrowding or other factors; growth and aging; and, finally, reproduction. For the hatchery oysters: (B) all parents are wild adults that will produce juvenile oysters, which are then planted into the Chesapeake Bay (orange arrow) and become part of the wild population. That will increase the wild population. Credit: Katie Hornick
This model will help us understand details of the genetic impacts of hatchery supplementation of oysters and will be useful for making decisions about which actions will be most effective within the Chesapeake Bay, as well as for planning newer restoration programs like that in the Hudson River in New York. This model will help to predict specific outcomes of management that we are currently unable to measure in the field. For example, what happens if we plant a lot of oysters with lower genetic diversity into the wild? Or what if one year there is a bad disease episode that wipes out a large proportion of the oyster population? Or, what if people engage in fishing at a site that is planted with hatchery oysters? Working to understand these specific impacts and integrating realistic problems into the model will provide useful information to managers and policy makers.
My hope is that this model can be used to better understand and improve our knowledge base in ways that will directly affect the collaborative oyster restoration efforts in the Chesapeake Bay. I especially hope that this model will be used to better to understand and/or inform new shellfish restoration projects around the world that will result in healthy, resilient, and self-sustaining populations of these species.
Photo, top: Image of a reef in Harris Creek. Photograph courtesy of Oyster Recovery Partnership
See all posts to the Fellowship Experiences blog




